Nearly 9% of residue side chains in the human proteome are primary amides belonging to asparagine, Asn, and glutamine, Gln, yet both residues have resisted chemoproteomic mapping at proteome scale. The obstacle is chemical inertness: unlike nucleophilic or redox-sensitive residues such as cysteine, lysine, and arginine, Asn and Gln amides carry no obvious handle for selective modification under mild aqueous conditions. That blind spot has left a substantial portion of proteome chemistry, including residues directly regulated by deamidation and N-glycosylation, largely uncharted and functionally underconnected.
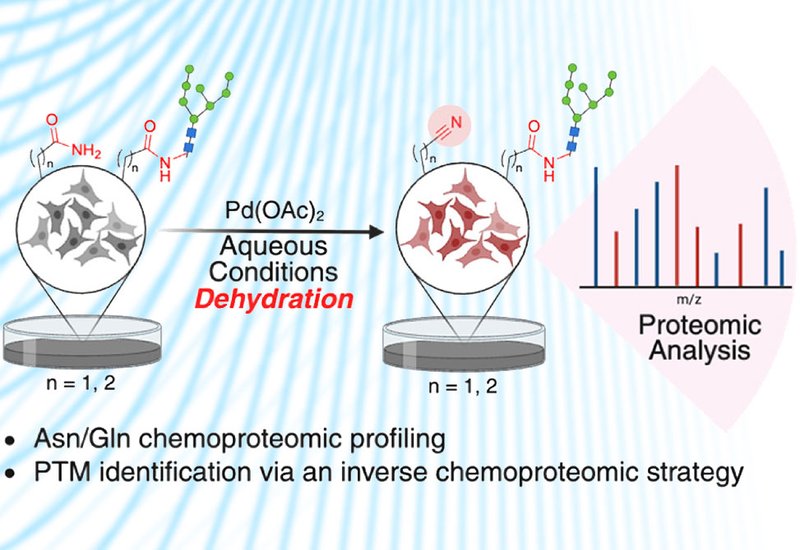
Researchers in the Raj Group at Emory University, published in ACS Chemical Biology, addresses that gap with a palladium-mediated dehydration strategy that converts Asn and Gln side-chain amides to nitriles through net loss of water under mild, aqueous conditions. The platform builds on a companion study from the same group, published in the Journal of the American Chemical Society, which established the mechanistic foundation for chemoselective Asn and Gln activation, including the palladium chelation strategy and the carbometalation scope that transforms the nitrile into a versatile bioorthogonal handle. In the chemoproteomic application reported here, the team used Pd(OAc)2 at low millimolar concentrations in a sodium phosphate buffer–acetonitrile mixture at room temperature, establishing chemoselective conversion of primary amides with no detectable cross-reactivity toward lysine, tyrosine, tryptophan, serine, methionine, histidine, or arginine. The nitrile product carries a net mass shift of –18.0106 Da, is chemically stable, and is readily detected by LC-MS/MS, providing a direct, label-free readout of chemically addressable Asn and Gln sites across complex proteomes. Nitrile-modified myoglobin retained peroxidase-like activity fully comparable to the unmodified protein, confirming that the chemistry operates without disrupting protein fold or function.
Applied to T-47D breast cancer cell lysates, dose-dependent treatment with Pd(OAc)2 identified 2,544 unique proteins at 1 mM, with 445 recurrent, high-occupancy conversion sites across 256 proteins spanning enzymes, regulatory factors, and structural proteins. Sequence-context analysis of hyperreactive sites revealed enrichment of nearby acidic and polar residues, pointing to local electrostatic and hydrogen-bonding environments as modulators of amide reactivity. Extension to intact T-47D cells at cytocompatible concentrations, with 92–96% viability across tested doses, identified 204 recurrent nitrile-bearing peptides across 141 proteins, with the live-cell dataset distinctly enriched for membrane-associated proteins including focal adhesion and anchoring junction components, a profile reflecting reagent access and subcellular compartmentalization rather than intrinsic amide reactivity alone. RNA-binding proteins emerged among the most consistently modified targets in both contexts, consistent with the known abundance of Asn and Gln in that functional class.
The team then deployed an inverse chemoproteomic logic: because post-translational modifications, PTMs, that convert or mask the amide, specifically deamidation to carboxylate and N-glycosylation of Asn, block nitrile formation, a protection signature directly reports PTM occupancy without requiring enrichment or enzymatic tagging. Under accelerated deamidation stress, nitrile-modified peptides dropped from 7,031 in untreated controls to 668 after 120 minutes, and sequence motif analysis of residues that remained nitrile-labeled recovered strong Asn-Pro enrichment, consistent with proline's known suppression of the succinimide intermediate that drives Asn deamidation. In Saccharomyces cerevisiae, tunicamycin-mediated inhibition of N-linked glycosylation produced the expected surge in nitrile-bearing peptides, 597 unique peptides compared with 163 in controls, and cross-referencing against the GlyCosmos database revealed that 86% of candidate glycoproteins identified carried no prior glycoprotein annotation, underscoring the complementarity of chemistry-based access to enrichment-centric workflows. Comparative profiling across yeast and hyphal states of the pathogen Candida albicans further captured morphogenesis-linked remodeling of N-glycosylation, nominating 34 proteins whose Asn nitrile signals were reduced or absent in the hyphal form, enriched in cell-wall organization, adhesion, and secretion pathways.
By converting a chemically inert functional group into a proteome-wide LC-MS readout, palladium-mediated amide dehydration broadens the residue-centric vocabulary of chemoproteomics to a previously inaccessible amino acid class. The platform complements existing nucleophile-targeted approaches and opens a direct experimental path toward connecting Asn and Gln chemistry, PTM dynamics, and protein function across physiology and disease, including host-pathogen biology where glycosylation remodeling is a central virulence mechanism.