Protein folding is governed by more than just the three-dimensional structure of the native state. The unfolded state, a heterogeneous ensemble of conformations that exists before folding or after denaturation, plays a critical role in determining stability and folding pathways. Yet the unfolded state remains difficult to study because it is sparsely populated under native conditions and highly dynamic. Traditional approaches to manipulate the unfolded state rely on side-chain mutagenesis, such as glycine substitutions to enhance flexibility or proline to restrict it. Backbone modification offers a complementary strategy: changing the conformational freedom of the polypeptide chain itself without altering side-chain identity. Despite this potential, rational application of backbone engineering to tune unfolded state properties in folded proteins has remained largely unexplored.
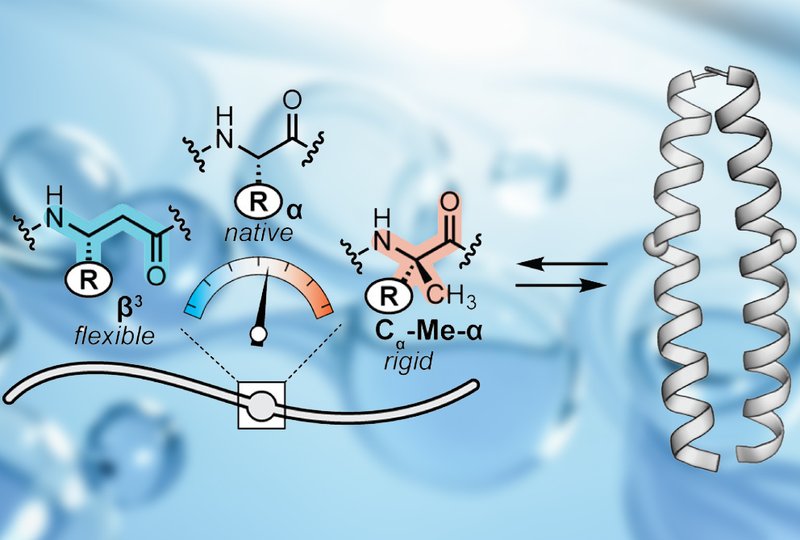
Researchers in the Horne Group at the University of Pittsburgh, published in Biochemistry, set out to develop a systematic method for manipulating the unfolded state of a well-folded protein through targeted backbone modification. They selected the GCN4 leucine zipper, a dimeric α-helical coiled coil, as their model system. The team designed 12 variants in which individual α-amino acids at solvent-exposed positions were replaced with either β3-residues or Cα-methylated α-residues, Cα-Me-α. These two modifications are regioisomeric, differing only in the placement of a single CH2 group. The β3-residue inserts CH2 between the carbonyl and α-carbon, adding a freely rotatable bond that enhances backbone flexibility. The Cα-Me-α residue inserts CH2 between the α-carbon and its hydrogen, creating geminal substitution that restricts conformational freedom. Crucially, both modifications retain the original side chain, isolating the effect to backbone dynamics. The variants were synthesized by solid-phase peptide synthesis and characterized by circular dichroism spectroscopy and X-ray crystallography.
X-ray crystal structures of six variants revealed that backbone modification had virtually no effect on the folded coiled coil structure. The variants superimposed on the prototype with backbone root-mean-square deviations of only 0.23 to 0.49 Å. Even salt bridges involving residues at modification sites remained intact. Circular dichroism spectra confirmed that all variants except one adopted α-helical secondary structures matching the prototype. Thermal stability varied widely depending on both the type and context of modification. Some variants were destabilized by more than 20 °C, while others gained up to 11 °C in melting temperature. Cα-Me-α substitutions were consistently more stabilizing than β3 replacements at the same position.
To probe thermodynamic parameters, the researchers performed coupled thermal and chemical denaturation experiments using urea. These measurements revealed complex context-dependent effects on folding free energy, enthalpy, and entropy. However, one parameter showed a strikingly uniform trend: the m value, which quantifies the sensitivity of folding free energy to denaturant concentration. The m value correlates with the change in solvent-accessible surface area between folded and unfolded states. Every β3 substitution increased the m value relative to the prototype, while every Cα-Me-α substitution decreased it. Because the folded structures are nearly identical, these changes in m value point directly to altered properties of the unfolded ensemble. Estimated solvent-accessible surface area changes varied by approximately twofold across the series. The team then tested dual-site variants combining two Cα-Me-α modifications. These showed even greater reductions in m value, demonstrating that mutually reinforcing modifications produce cumulative effects on the unfolded state.
The results establish backbone modification as a predictable tool for tuning unfolded state properties without changing side-chain composition or folded structure. The approach is orthogonal to traditional mutagenesis, offering researchers a new dimension for probing the role of conformational dynamics in protein folding. The findings have implications for fundamental studies of folding mechanisms, particularly investigations of transition state ensembles using kinetic analysis. The strategy is generalizable to other helical proteins accessible by chemical synthesis or semisynthesis. Future work combining backbone engineering with advanced biophysical methods such as NMR or single-molecule spectroscopy, or with molecular dynamics simulations, could provide unprecedented molecular-level insight into the structure and dynamics of unfolded protein ensembles.