Cancer cells evade immune surveillance through a layered set of adaptations: they shed or downregulate surface antigens, recruit immunosuppressive cells into the tumor microenvironment, TME, and drive T-cell exhaustion through persistent checkpoint signaling. Peptide-based vaccines aim to cut through these defenses by delivering tumor-specific antigen fragments directly to antigen-presenting cells, APCs, and training CD8+ and CD4+ T cells to seek and destroy malignant tissue. Compared to whole-cell or mRNA-based approaches, peptide vaccines offer lower manufacturing cost, longer circulation times, and straightforward conjugation to adjuvants or co-therapeutic payloads. Yet despite dozens of active clinical trials, not one peptide cancer vaccine has reached FDA approval, a gap that reflects unresolved problems in antigen selection, delivery, and overcoming the immunosuppressive TME.
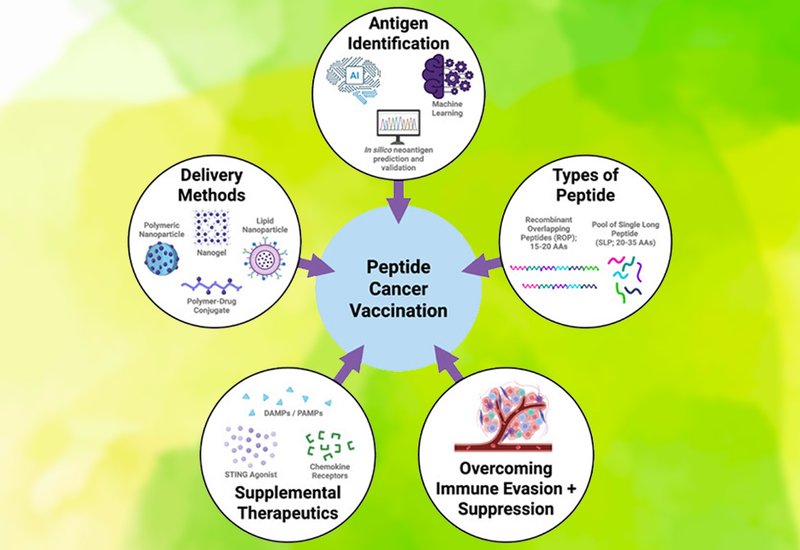
Researchers in the Kimmel Group at The Ohio State University, published in Biochemistry, surveyed the current state of peptide cancer vaccination across four interconnected challenges: identifying immunogenic neoantigens, formulating optimal peptide constructs, engineering delivery vehicles, and combining vaccines with adjuvants or checkpoint inhibitors. The review draws on findings from clinical trials, preclinical mouse models, and emerging computational platforms to map where the field stands and where the critical gaps remain.
A central theme is neoantigen identification. Tumor-specific antigens, TSAs, derived from somatic mutations offer higher target specificity and lower risk of off-target autoimmunity than tumor-associated antigens, TAAs, but are far harder to identify and validate. Historically, in silico sequencing predicted neoantigen candidates with limited clinical translation, because the predictions frequently miss peptides that fail to bind HLA molecules with sufficient affinity or that are rendered invisible by the immunosuppressive TME. Liquid chromatography-tandem mass spectrometry, LC-MS/MS, provides a direct readout of the HLA-presented ligandome but suffers from a bias toward abundant peptides under standard data-dependent acquisition. Manakongtreecheep et al. addressed this with Pepyrus, a workflow combining data-independent acquisition with machine learning to expand detectable neoantigen libraries. Complementing mass-spectrometry approaches, platforms such as TumorAgDB2.0 couple curated neoantigen databases with the NeoTImmuML prediction tool, which scores immunogenicity across 78 physicochemical features. Biochemical data reinforce these computational efforts: amino acid substitutions introducing large aromatic side chains correlate with stronger T-cell responses, and modifications to non-anchor residues that stabilize the peptide:MHC complex prolong TCR contact and sustain immunogenicity.
On the formulation side, synthetic long peptides, SLPs, of 25 to 35 residues outperform minimal epitopes by engaging both MHC class I and class II processing pathways, enabling CD4+/CD8+ co-activation. In a melanoma trial, SLP vaccines adjuvanted with a T-cell receptor agonist produced no disease progression in 67% of patients over 25 months, with persistent CD4+ and CD8+ responses in all evaluable participants. Recombinant overlapping peptides, ROPs, linked via the protease Cathepsin S, encode multiple epitopes in a single polypeptide chain and have generated protective cellular immunity in preclinical melanoma models; a first-in-human trial of OVM-200 is currently underway. For delivery, polymeric nanoparticles, lipid nanoparticles, cationic nanogels, and self-assembling peptide-TLR-7/8a conjugates each offer distinct advantages in protecting peptide cargo from enzymatic degradation, targeting lymph nodes, and sustaining APC uptake. Albumin-hitchhiking nanobody-STING agonist conjugates developed in the Kimmel group accumulated in tumors after systemic dosing, potentiated antitumor T-cell responses, and synergized with checkpoint blockade and adoptive cell therapy across multiple cancer subtypes. Combination with HER2-directed trastuzumab produced no disease recurrence over approximately 34 weeks in preliminary clinical data from HER2-positive breast cancer patients.
The review identifies three interlocking barriers to approval: the immunosuppressive TME, active immune evasion mechanisms including T-cell exhaustion and checkpoint upregulation, and tumor heterogeneity that enables antigen-loss escape variants. The authors argue that combination strategies pairing peptide vaccines with immune checkpoint inhibitors, STING agonists, or chemotherapy represent the most tractable near-term path to robust clinical efficacy. They also highlight that public neoantigens shared across patients, such as those arising from TP53 mutations, are more likely to be clonally expressed within tumors, reducing the risk of antigen-loss escape and making them practical targets for off-the-shelf formulations. Looking further ahead, the convergence of single-cell multi-omics, high-throughput yeast display screening, and AI-driven neoantigen discovery pipelines is expected to accelerate personalized vaccine design and extend peptide vaccination to rare cancers, infectious diseases including HIV and influenza, and settings where standard-of-care costs are prohibitive.