Quorum sensing allows bacteria to coordinate behavior across an entire population by secreting and detecting small signaling molecules. In Streptococcus sinensis, an oral commensal implicated in infective endocarditis, a severe cardiovascular disease with roughly 25% mortality, this communication depends on the competence stimulating peptide, CSP, which binds the membrane-bound histidine kinase receptor ComD and triggers expression of competence genes through the alternative sigma factor ComX. Disrupting this signaling circuit without killing the bacterium offers a path to anti-virulence therapy that sidesteps the selective pressure driving antibiotic resistance. Yet the molecular determinants of CSP recognition by ComD in S. sinensis had never been characterized.
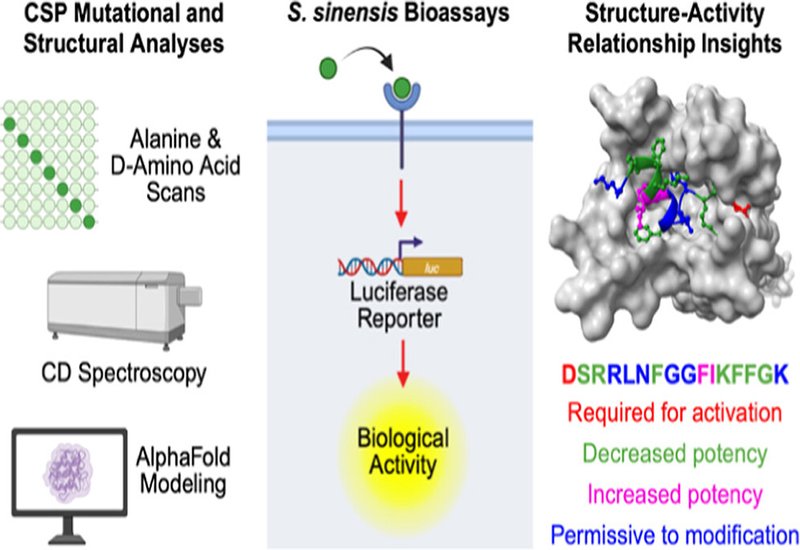
Researchers in the Tal-Gan Lab at the University of Nevada, Reno, published in Biochemistry, conducted the first comprehensive structure-activity relationship, SAR, analysis of the 16-residue S. sinensis CSP. To this end, the team prepared two complete analog libraries: one in which each residue was replaced sequentially with alanine, and one in which each residue was replaced with its D-configured counterpart. To evaluate receptor activation without interference from endogenous peptide, the group engineered a dedicated reporter strain, S. sinensis ΔcomC PcomX-luc, in which the comC gene was knocked out and a firefly luciferase reporter placed under control of the comX promoter, providing a quantitative readout of ComD activity.
Agonism screening identified most single-residue alanine substitutions as receptor agonists, with EC50 values spanning 2.14 to 348 nM compared with 8.72 nM for the native CSP. Substitutions at F10 and I11 increased potency, with CSP-I11A reaching an EC50 of 2.14 nM, approximately four-fold more potent than native CSP. Positions R3, F7, K12, F13, F14, and G15 were important for efficient receptor activation, as their alanine analogs required higher concentrations for full activation. Notably, the S. sinensis ComD receptor was more permissive to sequence variation than its counterparts in related Mitis-group streptococci: even CSP-R3A, the weakest agonist in the alanine scan, retained biologically relevant activity, a marked contrast to the near-complete loss of binding observed at the equivalent position in S. pneumoniae, S. mitis, and several other streptococcal species.
The single exception to agonism was the N-terminal substitution CSP-D1A, which converted the native activating signal into a competitive inhibitor with an IC50 of 293 nM. D-amino acid scanning reinforced this picture: replacing D-Asp at position 1 with its D-configured counterpart preserved full agonist activity with an EC50 of 9.51 nM, indicating that negative charge rather than side-chain geometry drives receptor activation at this site. By contrast, D-amino acid substitutions at most internal positions reduced potency substantially, with CSP-i11 reaching an EC50 of 348 nM, the weakest D-analog, while CSP-I11A was the strongest agonist overall. The inversion at that single position produced opposing functional outcomes, underscoring the spatial specificity of the receptor binding pocket in that region. Circular dichroism spectroscopy in membrane-mimicking conditions revealed that native CSP and most analogs adopt only weak or transient α-helical structure, with 6.6% helicity for the native peptide, cautioning against assignment of a rigid helical binding interface. AlphaFold 3 Multimer modeling placed CSP in a partial α-helical conformation contacting the transmembrane region adjoining the DHp domain of ComD, consistent with the SAR requirement for an intact N-terminus. Superimposition of the CSP/ComD and CSP-D1A/ComD models yielded an RMSD of 1.337, without major differences in binding orientation or receptor conformation, illustrating the limits of static structure prediction for resolving activation dynamics.
This work delivers the first molecular blueprint of S. sinensis CSP signaling and identifies CSP-D1A as a lead competitive inhibitor of the competence regulon. The relatively permissive ComD binding interface, combined with discrete hotspot residues at D1, R3, F7, and the aromatic cluster F13–F14, provides a rational foundation for optimizing inhibitory analogs. The authors note that future truncation scanning and combinatorial library development should define the minimal scaffold for potent ComD binding, and that phenotypic validation of CSP-D1A through biofilm formation assays and pathogenesis models will be necessary to establish translational impact. Incorporating D-amino acids at permissive positions such as D1, S2, K12, and K16 could improve proteolytic stability without sacrificing inhibitory potency, supporting the broader goal of developing anti-virulence therapeutics against S. sinensis-associated infective endocarditis.